As the longest-standing Neuroscience PhD program in the country, we have a history of training leaders in the field. Join our multidisciplinary program and begin to forge your future of excellence.

The country’s first of its kind, U-M Medical School’s Neuroscience training program began in 1971 and continues to lead the industry today.
Our difference is rooted in our interdisciplinary and interdepartmental approach, with more than 150 core affiliated faculty members distributed throughout our institution. We are a collegial and interactive group that performs research across the breadth of the neuroscience field.
Our faculty includes members of the National Academy of Sciences, National Academy of Medicine, past presidents of the Society for Neuroscience, fellows of the AAAS and Highly Cited Researchers in their fields.
204 Washtenaw Ave.
Ann Arbor, Michigan 48109-2215
Watch and learn what it’s like to immerse yourself in the Neuroscience Graduate Program at the University of Michigan Medical School.

Your partnership will support our world-class, neuroscience graduate student training. Explore opportunities to support our important work and further our goals.

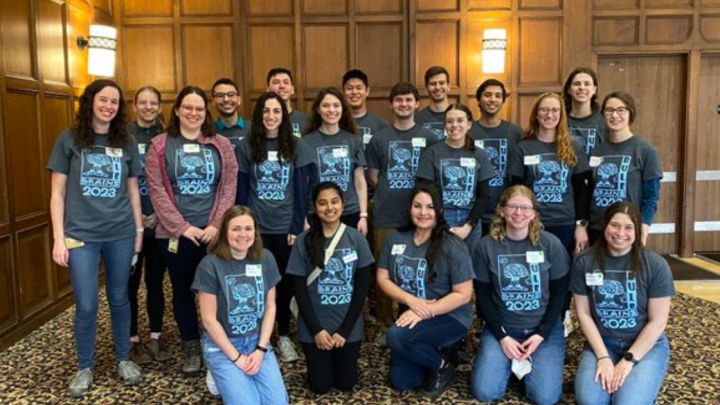
Our graduate program is committed to diversity, equity and inclusion with comprehensive opportunities to get involved and commit to making lasting change.
The Neuroscience Graduate Program’s ultimate goal is to prepare the future leaders in the field of neuroscience by providing the training and expertise necessary to succeed in any scientific career the students may choose. Welcome and Go Blue!”